 Request a FREE Sample of our FcRn Binding Kit!
Request a FREE Sample of our FcRn Binding Kit! Request a FREE Sample of our Fc gamma RI / CD64 Binding Kit !
Request a FREE Sample of our Fc gamma RI / CD64 Binding Kit !
 Happy Holiday! Limited Keychain here with your next order
Happy Holiday! Limited Keychain here with your next order Happy Holiday! Limited Keychain here with your next order
Happy Holiday! Limited Keychain here with your next order
 Request a FREE sample of our GMP products!
Request a FREE sample of our GMP products!  Request a FREE sample of our GMP products!
Request a FREE sample of our GMP products!
The emergence of SARS-CoV-2 has led to over 1.8 million infections and over 118 thousand deaths. The number is still growing at this moment. Similar to SARS-CoV and MERS-CoV, SARS-CoV-2 is highly transmissible from infected individuals, even without symptoms, to healthy ones, which can cause extremely severe respiratory symptoms. The need to rapidly develop vaccines as well as therapeutics against SARS-CoV-2 at a time of the explosion is imperative.
Coronavirus entry into the host cell is mediated through the transmembrane spike (S) glycoprotein that forms homotrimers protruding from the viral surface. This S glycoprotein is the main target of neutralizing antibodies upon infection as well as the focus of therapeutic and vaccine design. S trimers are extensively decorated with N-linked glycans that are important for proper folding and for modulating accessibility to host proteases.
Here in this article, we would like to walk you through a few recent studies to explore more scientific details about the glycosylation of the S protein and the importance of it.
1. Site-specific analysis of the SARS-CoV-2 glycan shield[1]
Crispin team successfully expressed and purified recombinant SARS-CoV-2 S protein from HEK293F cells. They studied the composition of site-specific N-linked glycan as well as the degree of sequon occupancy on the protein using a mass spectrometer. All 22 glycans on the SARS-CoV-2 S protein were identified, and 18 of those N-glycosites were conserved between SARS-CoV and SARS-CoV-2 S proteins (Figure 1).
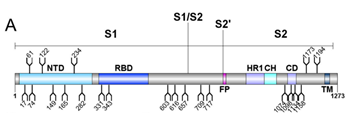
Figure 1. Schematic representation of SARS-CoV-2 S glycoprotein.
The abundances of each glycan are summed into oligomannose-, hybrid, and complex-type glycosylation, revealing the extensive heterogeneity in N-glycan composition and site occupancy. Besides, the diverse signals arising from heterogeneous complex-type glycosylation are simplified by the summation of glycan intensities into a more limited range of structural categories (Figure 2).
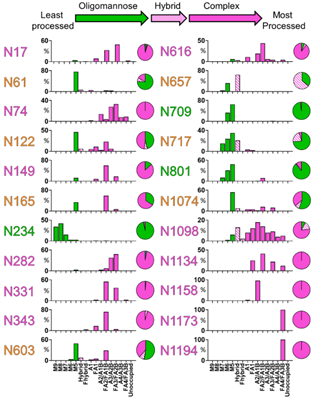
Figure 2. Site-specific N-linked glycosylation of SARS-CoV-2 S glycoprotein.
2. Site-specific N-glycosylation Characterization of Recombinant SARS-CoV-2 Spike Proteins using High-Resolution Mass Spectrometry[2]
A team from West China Hospital of Sichuan University published their work recently. They characterized the site-specific N-glycosylation of HEK293 cell expressed SARS-CoV-2 S1 protein and insect cell expressed S protein. Following digestion with two complementary proteases to cover all potential N-glycosylation sequons and integrated N-glycoproteomics analysis, they performed stepped collision energy (SCE) mass spectrometry to reveal the N-glycosylation profile of SARS-CoV-2 S proteins at the levels of intact N-glycopeptides and glycosites, along with the glycan composition and site-specific number of glycans (Figure 3).
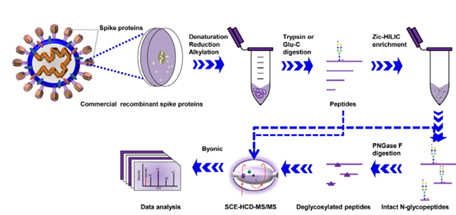
Figure 3. Workflow for site-specific N-glycosylation characterization of recombinant SARS-CoV-2 S proteins using two complementary proteases for digestion and simultaneous N-glycoproteomics analysis.
All 22 potential canonical N-glycosites were identified in S protein protomer. Notably, insect cell-produced SARS-CoV-2 S protein contained 38 N-glycans, which were mainly high-mannose type N-glycans, whereas the human cell-produced protein possessed up to 140 N-glycans, most of which are complex type (Figure 4). This N-glycosylation profiling and determination of differences between distinct expression systems could shed light on the infection mechanism and promote the development of vaccines and targeted therapeutics.
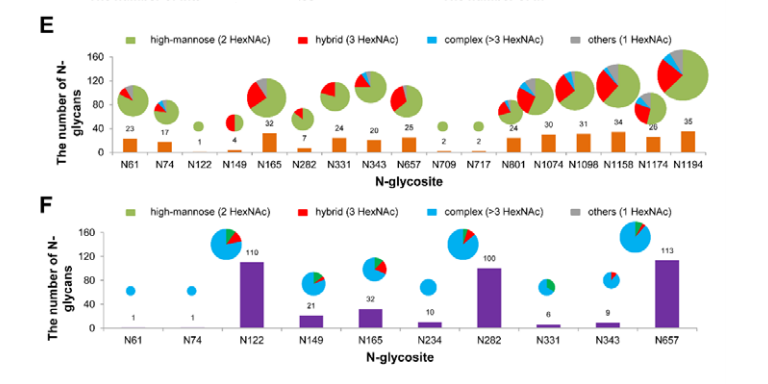
Figure 4. Site-specific N-glycosylation of recombinant SARS-CoV-2 S proteins.
3. Deducing the N- and O- glycosylation profile of the spike protein of novel coronavirus SARS-CoV-2[3]
Azadi’s team from the University of Georgia published their work on bioRxiv. They digested the recombinant SARS-CoV-2 subunit S1 and S2 expressed in HEK293 cells into the glycopeptides. After that, high resolution LC-MS/MS was used to analyze these glycopeptides. In this way, they identified partial N-glycan occupancy on 17 out of 22 N-glycosylation sites with a combination of high mannose and complex type, but no hybrid-type glycans (Figure 5). Interestingly, they observed two unexpected O-glycosylation modifications on the receptor binding domain (RBD) of spike protein subunit S1. Even though O-glycosylation has been predicted on the spike protein of SARS-Cov-2, this is the first report of the site of O-glycosylation and identity of the O-glycans attached on the subunit S1.
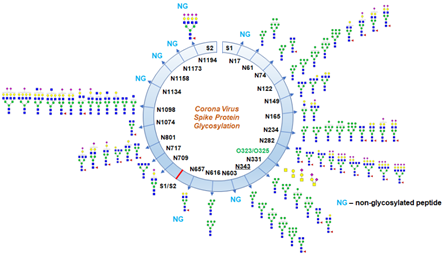
Figure 5. Glycosylation profile on coronavirus SARS-CoV-2 characterized by high-resolution LCMS/MS.
4. Structural and functional analysis of a potent sarbecovirus neutralizing antibody [4]
Vir Biotechnology collaborated with the University of Washington to study the potential cross reactivity between SARS-CoV-2 and the neutralizing monoclonal antibodies identified from an individual infected with SARS-CoV in 2003. mAb S309 potently neutralized SARS-CoV-2, SARS-CoV pseudoviruses as well as the authentic SARS-CoV-2 by engaging the S receptor binding domain (Figure 6).
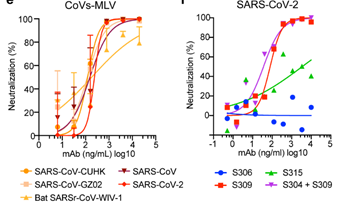
Figure 6. Neutralization of pseudoviruses (bearing S protein) and authentic SARS-CoV-2 by an isolated antibody named S309 from a SARS survivor
It was shown through cryo-electron microscopic analysis and different binding assays that S309 recognized a glycan-containing epitope, which is conserved within the sarbecovirus subgenus, without competing with receptor binding (Figure 7). The glycan recognition of S309 implies the importance of the naturally decorated N-glycans in SARS-CoV-2 S proteins.
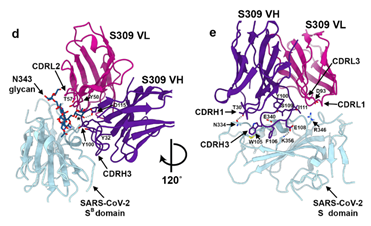
Figure 7. Closeup view of the S309 epitope showing the contacts formed with the core fucose (labeled with a star) and the core N-acetyl-glucosamine of the oligosaccharide at position N343.
5. 3D Models of glycosylated SARS-CoV-2 spike protein suggest challenges and opportunities for vaccine development [5]
A group of researchers from the University of Georgia reported their work about the 3D models of glycosylated SARS-CoV-2 spike protein. Their molecular dynamics simulation result revealed that glycans shield approximately 40% of the underlying protein surface of the S glycoprotein from antibody recognition, but the efficacy of antisera should not be impacted. The most accessible and largest antigenic surface in the S protein consists of the ACE2 binding domain, suggesting that a vaccine with this epitope may be effective, as long as the virus continues to target the same host receptor (Figure 8). Furthermore, efficacy may benefit from ensuring that the production method results in glycosylation profiles that match those of the circulating virus, or by engineering a vaccine to avoid glycosylated sequences.
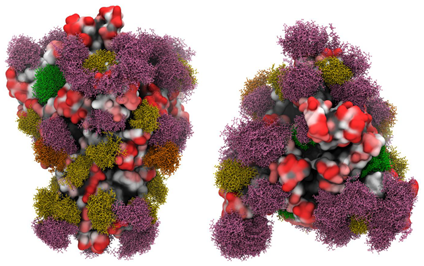
Figure 8. Side view (left panels) and Top view (right panels) of the SARS-CoV-2 S glycoprotein with site-specific glycosylation.
As we can see from the previous studies, glycosylation of the S protein is complex and important as well. Different expressing systems can produce proteins with totally different glycosylation results in terms of the composition and type of glycan. Importantly, the glycosylation can affect the vaccine and therapeutics development, which should draw much more attention from scientists worldwide. The drug industry is hoping to compress the time it takes to get a vaccine and the therapeutics to market — usually about 10 to 15 years — to within the next year. Wisely choose the protein reagent will help to save a lot of troubles.
Reference:
1. Yasunori Watanabe et al. Site-specific analysis of the SARS-CoV-2 glycan shield. bioRxiv preprint doi: https://doi.org/10.1101/2020.03.26.010322
2. Yong Zhang et al. Site-specific N-glycosylation Characterization of Recombinant SARS-CoV-2 Spike Proteins using High-Resolution Mass Spectrometry. bioRxiv preprint doi: https://doi.org/10.1101/2020.03.28.013276.
3. Asif Shajahan et al. Deducing the N- and O- glycosylation profile of the spike protein of novel coronavirus SARS-CoV-2. bioRxiv preprint doi: https://doi.org/10.1101/2020.04.01.020966.
4. Dora Pinto et al. Structural and functional analysis of a potent sarbecovirus neutralizing antibody. bioRxiv preprint doi: https://doi.org/10.1101/2020.04.07.023903.
5. Oliver C. Grant et al. 3D Models of glycosylated SARS-CoV-2 spike protein suggest challenges and opportunities for vaccine development. bioRxiv preprint doi: http://biorxiv.org/cgi/content/short/2020.04.07.030445.
6. https://www.who.int/emergencies/diseases/novel-coronavirus-2019/situation-reports/
7. Walls et al., Structure, Function, and Antigenicity of the SARS-CoV-2 Spike Glycoprotein, Cell (2020).
8. Walls, A.C. et al. Glycan shield and epitope masking of a coronavirus spike protein observed by cryo-electron microscopy. Nat. Struct. Mol. Biol. 23, 899–905. (2016).
9. Walls, A.C. et al. Tectonic conformational changes of a coronavirus spike glycoprotein promote membrane fusion. Proc. Natl. Acad. Sci. USA 114, 11157–11162. (2017).
ACROBiosystems has developed various SARS-CoV-2 proteins based on their HEK293 protein expression platform. The state of their mammalian cell expressed recombinant proteins is closer to that of the protein in human including the glycosylation level.
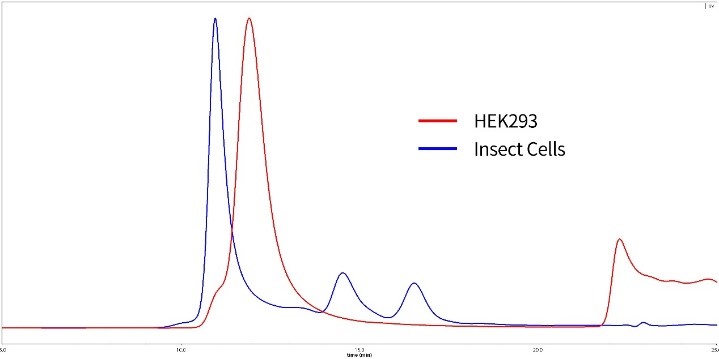
Figure 9. SEC-MALS of HEK293 expressed and insect cell expressed S proteins
Red: HEK293 expressed S protein is verified to be trimer with 85% purity. MW is 600 to 660kDa.
Blue: Insect cell expressed S protein is in high polymerization state.
(Related ACRO product: SARS-CoV-2 (COVID-19) S protein (R683A, R685A), His Tag, active trimer, HEK 293)
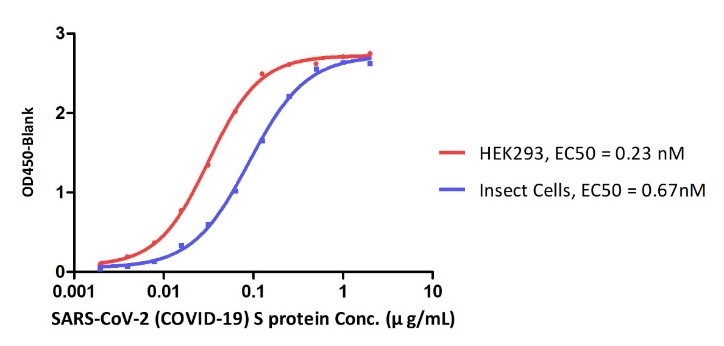
Figure 10. ELISA Binding affinity between ACE2 protein and S proteins expressed by HEK293 cell and insect cell respectively.
Their data revealed that SARS-CoV-2 S proteins expressed by different expressing systems can result in the different binding affinities to ACE2. This could due to the different post translational modifications especially the glycosylation between two different expressing systems.
Apart from the S protein, ACROBiosystems developed S, S1, S-RBD and ACE2 proteins, which are verified by Biacore and ELISA assays. Intended for the development of the vaccine, therapeutics and diagnostic cassettes, these HEK293 expressed SARS-CoV-2 proteins do have advantages in terms of mimicking the condition in blood circulation. Please check product details below :
This web search service is supported by Google Inc.
