 Request a FREE Sample of our FcRn Binding Kit!
Request a FREE Sample of our FcRn Binding Kit! Request a FREE Sample of our FcRn Binding Kit!
Request a FREE Sample of our FcRn Binding Kit!
 Limited Edition Golden Llama is here! Check out how you can get one.
Limited Edition Golden Llama is here! Check out how you can get one.  Limited Edition Golden Llama is here! Check out how you can get one.
Limited Edition Golden Llama is here! Check out how you can get one.
 Request a FREE sample of our GMP products!
Request a FREE sample of our GMP products!  Request a FREE sample of our GMP products!
Request a FREE sample of our GMP products!
> Insights > 【Inspiring Target】BiKEs and TriKEs, specifically targeting CD16a, showing new prospects for cancer therapy Natural killer (NK) cells are important effector cells of the innate immune system, which can quickly recognize and eliminate infection, stress, and malignant cells. Unlike T cells and B cells expressing antigen-recognition receptors such as TCR and BCR, NK cells can destroy target cells without prior antigen sensitization (e.g., virus-infected cells, certain tumor cells, and damaged cells). In addition, NK cells have no MHC restriction and fast response speed, which can play a role in the early stage of the immune response. This is also an important reason why NK cells are called natural killer cells.
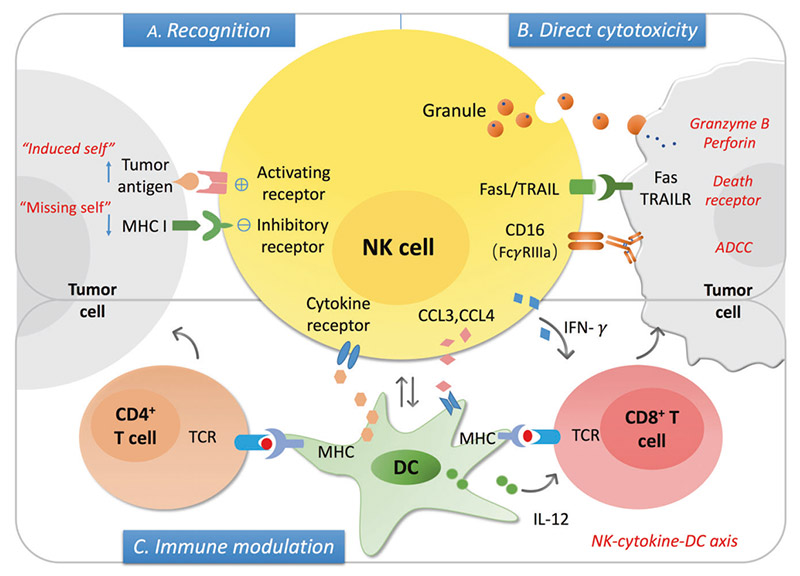
The mechanism of NK cells in cancer
NK cells can be used as innate cytotoxic effectors to regulate innate and adaptive immunity. The levels of stress-induced ligand (" induced self ") and MHC I decreased expression (" miss self ") can be recognized by activating and inhibiting receptors on NK cells respectively, and the balance between the two determines the activation state of NK cells. After activation, NK cells can induce the lysis of the target cells via secreting granules with granzyme B and perforin, the death receptor/ligand interaction, and antibody-dependent cell cytotoxicity (ADCC). On the other hand, activation of NK cells secretes a series of cytokines and chemokines that promote DC maturation and recruitment, further regulating T cell responses, including cytotoxic CD8+T cells and Th cell activation.
ADCC is a process involved in NK cell activation, release of cytokines and cytolytic particles, and induction of apoptosis of target cells, mainly mediated by low-affinity Fc receptor, FcγRIIIA/CD16a. CD16 is a receptor that activates the NK cell effect, inducing tyrosine-based activation motif (ITAM) phosphorylation of the immune receptor through signal transduction, triggering the release of dissolved particles and cytokines such as IFN-γ and TNF-α. In addition, when NK cells interact with target cells, CD16 in NK cells is down-regulated, which results from the shedding of CD16 exodomains during NK cell activation. CD16 shedding of NK cells is mediated by transmembrane glycoprotein, collapse, and metalloproteinase 17 or ADAM17. All of these demonstrate the central importance of CD16 in triggering NK cell-mediated ADCC.
Innovative techniques using NK cells in immunotherapy have introduced bi-specific killer cell engagers (BiKEs) and Tri-specific Killer cell Engagers (TriKEs). BiKEs were created from the fusion of anti-CD16 scFv and anti-tumor-associated antigen scFv. TriKEs are scFv combinations that anti-CD16 and two tumor-associated antigens. These molecules directly activate NK cells through CD16, amplify NK cells, and produce cytokines that have cytotoxic activity against various tumor cell antigens. TriKEs are usually produced by variable domain genes of specific monoclonal antibodies. In this case, antigen-binding site 1 is bivalent and engages with an antigen on effector cell, antigen-binding sites 2 and 3 engage antigens 1 and 2 on tumor cells.
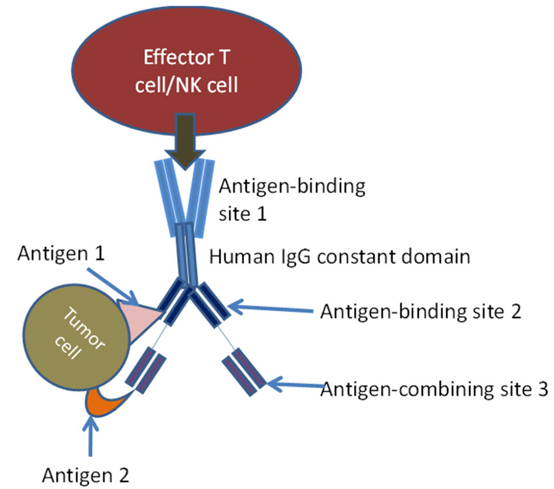
Schematic diagram of tri-specific antibody
Multiple BiKEs and TriKEs are currently in preclinical studies to directly activate NK cells, promote the formation of immune synapses between NK cells and tumor cells, and have cytotoxic activity against various tumor cell antigens through CD16.
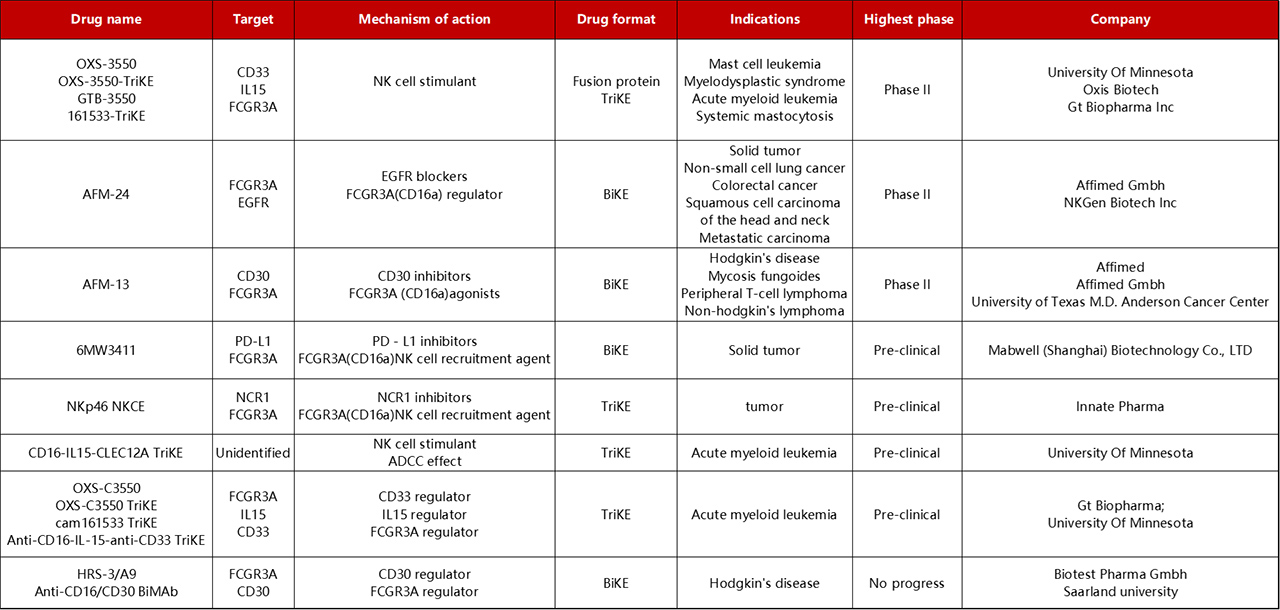
Progress of BiKEs and TriKEs in drug development
AFM13 is first-in-class tetravalent BiKEs developed by Affimed. One arm of AFM13 binds to the CD30 antigen on lymphoma cells, whereas the other arm binds to the CD16A antigen on the NK cells. The activated NK cells destroy the lymphoma cells. The NK cell activation and lymphoma destruction mediated by AFM13 are CD30-dependent.
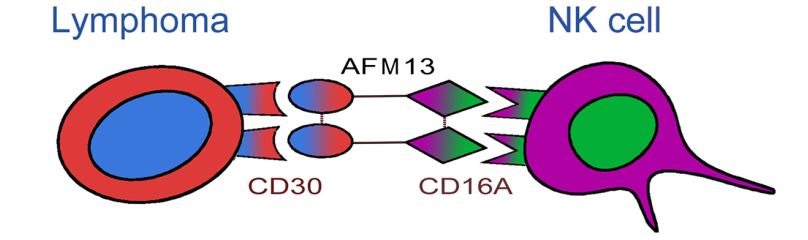
Structure of AFM13
Binding of AFM13 to CD30+ malignancies enhance NK cell activation, cytotoxicity and secretion of cytokines and chemokines, and AFM13 has shown good efficacy data in both monotherapy and combination therapy with anti-PD-1 for CD30+ malignancies.
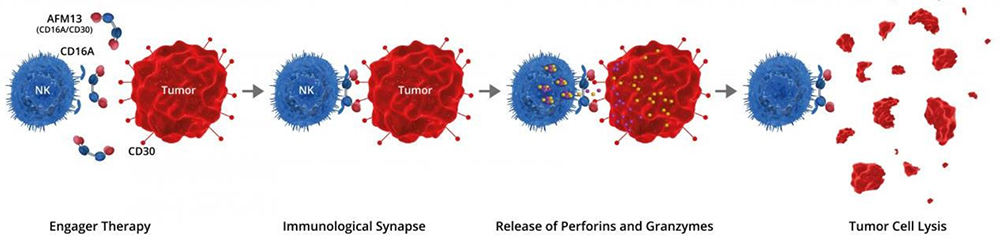
AFM13-mediated activation
AFM24, also from Affimed, is a new a novel bispecifc, IgG1-scFv fusion antibody targeting CD16a on innate immune cells, and EGFR on tumor cells using the redirected optimized cell killing (ROCK®) antibody platform. Studies have shown that AFM24 binds CD16a on NK cells and macrophages in the low nanomolar range, and binds to a variety of EGFR-expressing tumor cells. AFM24 is highly effective in vitro through NK cell-mediated cytotoxicity of antibody-dependent cells and macrophage-mediated ADCP. More importantly, AFM24 was effective against multiple EGFR-expressing tumor cells, targeting a variety of solid tumors, such as colorectal cancer or lung cancer, regardless of EGFR expression level and KRAS/ BRAF mutation. In vivo data showed that cynomolgus tolerated AFM24 at the highest dose (75 mg/kg) once a week for 28 days, and no dermal or other toxicity was observed. A temporary increase in IL-6 detected at all doses 2-4 h later returned to baseline levels after 24 h. These results demonstrate the potential of AFM24 to effectively target tumors expressing different levels of EGFR, regardless of their mutation, and highlight the promise of BiKEs as an alternative cancer therapy.
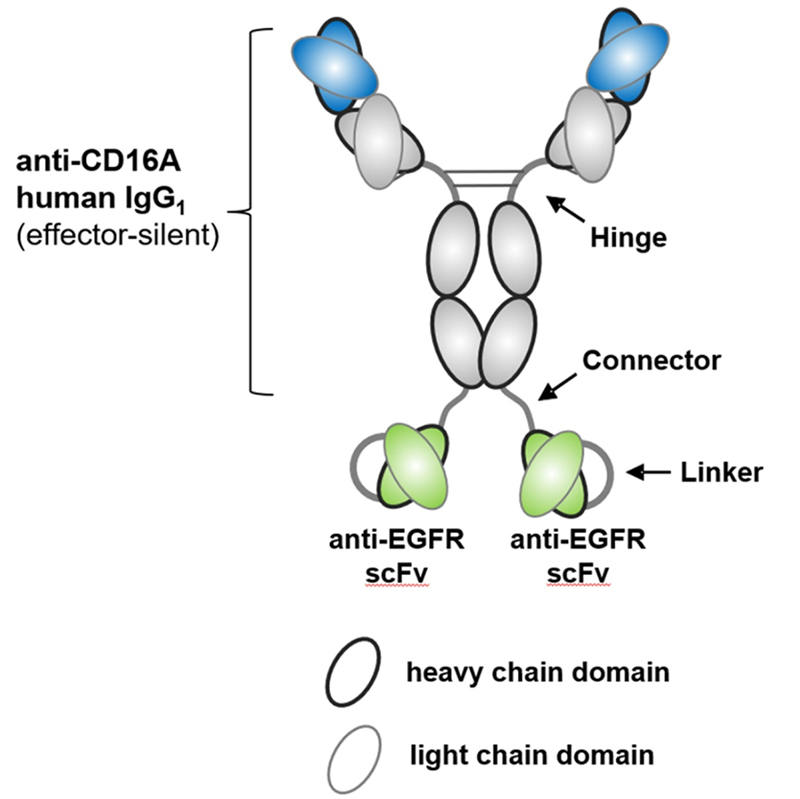
Structure of AFM24
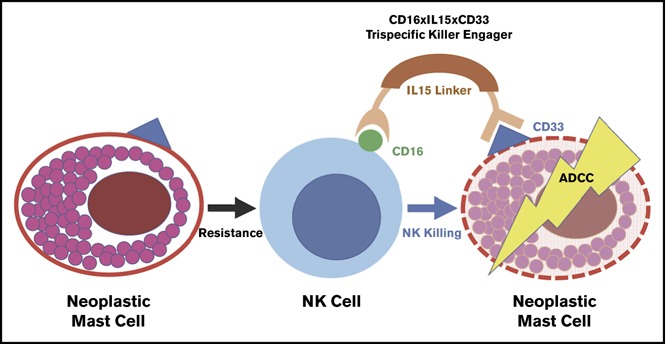
Structure of 61533
TriKE 61533 combines anti-CD16 scFv with anti-CD33 scFv, IL-15 is inserted as a linker between them to amplify NK cells and enhance their mediated killing of CD33- targeted myeloid cell. Described as a first-generation TriKE, triKE 161533 (SCFV16-M15-33) contains a mutant IL-15 with an N72D substitution (abbreviated m15), and a phase I/II clinical trial is underway for the treatment of refractory AML, high-risk MDS, and systemic mastocytosis (NCT03214666).
Further studies have shown that the effect of substituting a “humanized” anti-CD16 single domain, camelid antibody to replace the antiCD16 scFv, which enabled the presentation of wild-type (wt) IL15 without the use of m15. The resulting molecule, cam16-wt15–33 TriKE, displays stronger IL15 signaling capabilities, better NK cell activation, and increased NK cell–mediated tumor control both in vitro and invivo.
The TriKE platform is unique among NK engagers because it incorporates cytokine signaling, tumor-specific targeting, and activation of a single intramolecular ADCC. In summary, TriKEs not only drive specific tumor-killing, but also induce NK cell proliferation and survival via IL-15 fragments.
Although most BiKEs and TriKEs are currently in the preclinical stage, their ability to enhance NK cell-mediated cytotoxicity against a target holds great promise in treating cancer patients. Further investigation is needed to evaluate the safety and efficacy of BiKE and TriKE before clinical use.
CD16 (FcγRIII) is a low-affinity receptor for the IgG Fc domain and Human CD16 has two isoforms, CD16a and CD16b, their immunoglobulin binding regions share 96% of the same sequence. CD16a is an activating receptor mainly expressed on NK cells and macrophages. CD16b is expressed mainly on granulocytes and is not involved in tumor cell killing. At present, most antibodies targeting CD16 cannot distinguish between CD16a and CD16b. Considering CD16b cannot cause a killing effect on tumor cells and may cause side effects. Therefore, specific targeting of CD16a is particularly important, which is also one of the core technologies of Affimed.
ACROBiosystems has successfully developed a range of high-quality Fc receptor proteins to support antibody drug development and enable functional validation of the Fc domain of antibodies. Human FcγRIIIA/CD16a and its 96% homology FcγRIIIB/CD16b (NA1 and NA2), Mouse, Cynomolgus, Rhesus Macaque FcγRIII/CD16, Mouse FcγRIV/CD16-2 are available to accelerate the research and drug development that targeting FcγRIIIA/CD16a.
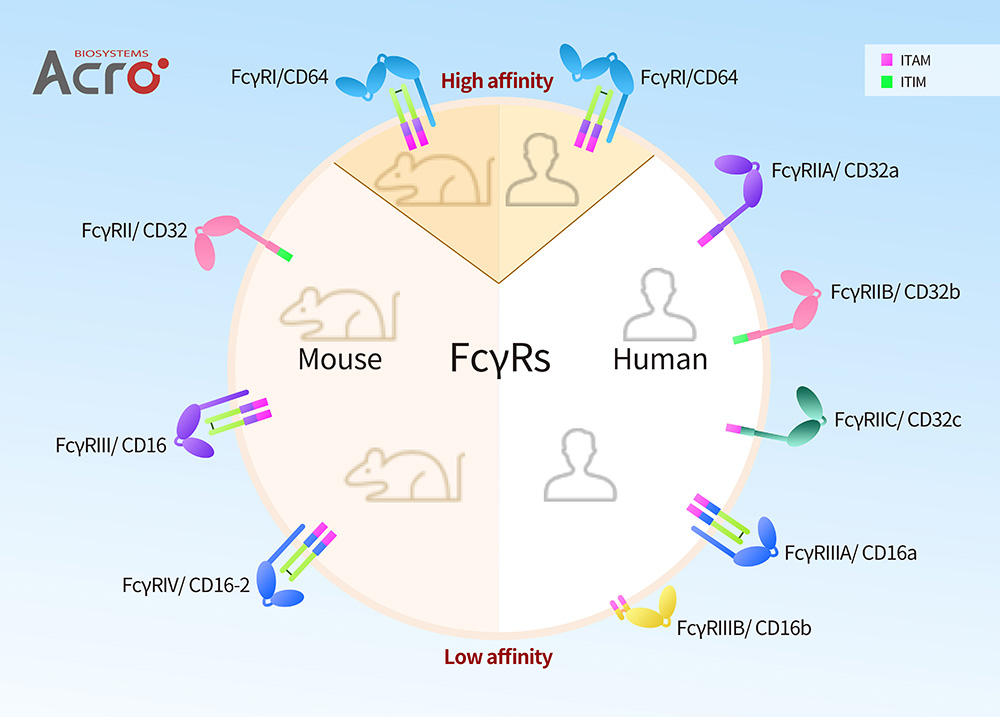
FcγRs in Human and Mouse
As is shown in the above figure, Human FcγRs can be classified FcγRI/CD64, FcγRII/CD32 (FcγRIIa/CD32a, FcγRIIb/CD32b and FcγRIIc/CD32c) and FcγRIII/CD16 (FcγRIIIA/ CD16a and FcγRIIIB/CD16b). Mouse FcγRs are classified as FcγRI/CD64, FcγRII/CD32, FcγRIII/CD16, and FcγRIV/CD16-2, which can be described as the equivalent of Human FcγRI/CD64, FcγRIIB/CD32b, FcγRIIA/CD32a, and FcγRIIIA/CD16a respectively according to homologues, including comparative binding information, significant polymorphisms, and important domains. FcγRI/CD64, FcγRIII/CD16, FcγRIV/CD16-2 showed activation characteristics corresponding to human, and FcγRII showed the same inhibitory function as human FcγRIIB.
 Expressed by HEK293 Cells: post-translational modification and proper protein folding
Expressed by HEK293 Cells: post-translational modification and proper protein folding
 Multiple species:Human FcγRIIIA/CD16a and FcγRIIIB/CD16b,Mouse, Cynomolgus, Rhesus macaque FcγRIII/CD16,Mouse FcγRIV/CD16-2 are available and can be used to detect the affinity between antibody candidates and CD16 molecule.
Multiple species:Human FcγRIIIA/CD16a and FcγRIIIB/CD16b,Mouse, Cynomolgus, Rhesus macaque FcγRIII/CD16,Mouse FcγRIV/CD16-2 are available and can be used to detect the affinity between antibody candidates and CD16 molecule.
 Natural mutants included (mutants containing signal peptide sequence count)(The sequence counting of mutation sites is containing signal peptides):Considering the mutation polymorphism of Human Fc receptors. We have developed two forms of Human CD16a, V176 and F176. And Human CD16b includes two forms, NA1 and NA2, the NA1 form of CD16b differs from the NA2 form of CD16b in AA36, 65, 82, and 106. The NA1 form of CD16b contains R36, N65, D82, and V106, while the NA2 form of CD16b contains S36, S65, N82, and I106.
Natural mutants included (mutants containing signal peptide sequence count)(The sequence counting of mutation sites is containing signal peptides):Considering the mutation polymorphism of Human Fc receptors. We have developed two forms of Human CD16a, V176 and F176. And Human CD16b includes two forms, NA1 and NA2, the NA1 form of CD16b differs from the NA2 form of CD16b in AA36, 65, 82, and 106. The NA1 form of CD16b contains R36, N65, D82, and V106, while the NA2 form of CD16b contains S36, S65, N82, and I106.
 Biotinylated CD16 proteins labeled with AvitagTM offered: the labeling efficiency is high, and the labeling site is specific and clear, which is suitable for ELISA/SPR/BLI detection based on binding to streptavidin in the process of drug development and optimization process
Biotinylated CD16 proteins labeled with AvitagTM offered: the labeling efficiency is high, and the labeling site is specific and clear, which is suitable for ELISA/SPR/BLI detection based on binding to streptavidin in the process of drug development and optimization process
 High purity: SDS-PAGE verification purity>95%, SEC-MALS verification purity>90%(QC standard)
High purity: SDS-PAGE verification purity>95%, SEC-MALS verification purity>90%(QC standard)
 Low endotoxin: <1.0 EU/µg
Low endotoxin: <1.0 EU/µg
 High stability: strict quality control to ensure high batch-to-batch consistency
High stability: strict quality control to ensure high batch-to-batch consistency
 Affinity verified by SPR & BLI: activity guaranteed, and protocols offered for free
Affinity verified by SPR & BLI: activity guaranteed, and protocols offered for free
CD16
| Molecule | Cat. No. | Host | Product Description | Structure |
|---|---|---|---|---|
| Fc gamma RIIIA / CD16a | CD8-H52H4 | HEK293 | Human Fc gamma RIIIA / CD16a (V176) Protein, His Tag (SPR & BLI & MALS verified) |  |
| CDA-H5220 | HEK293 | Human Fc gamma RIIIA / CD16a (F176) Protein, His Tag (SPR & BLI & MALS verified) |  | |
| CDA-H5290 | HEK293 | Human Fc gamma RIIIA / CD16a (V176) Protein, SUMO,His Tag (MALS & BLI verified) |  | |
| CDA-H52H6 | HEK293 | Human Fc gamma RIIIA / CD16a (F176, S197P) Protein, His Tag (MALS & SPR verified) |  | |
| CDA-H52S1 | HEK293 | Human Fc gamma RIIIA / CD16a (V176) Protein, HSA,His Tag (MALS & BLI verified) |  | |
| CDA-H82E8 | HEK293 | Biotinylated Human Fc gamma RIIIA / CD16a (F176) Protein,,His Tag (SPR & BLI & MALS verified) |  | |
| CDA-H82E9 | HEK293 | Biotinylated Human Fc gamma RIIIA / CD16a (V176) Protein, ,His Tag (SPR & BLI verified) |  | |
| Fc gamma RIIIB / CD16b (NA1) | CDB-H5227 | HEK293 | Human Fc gamma RIIIB / CD16b (NA1) Protein, His Tag (SPR & BLI & MALS verified) |  |
| CDB-H5296 | HEK293 | Human Fc gamma RIIIB / CD16b (NA1) Protein, SUMO,His Tag (MALS & BLI-verified) |  | |
| CDB-H82E4 | HEK293 | Biotinylated Human Fc gamma RIIIB / CD16b (NA1) Protein, His,(SPR & BLI & MALS verified) |  | |
| Fc gamma RIIIB / CD16b (NA2) | CDB-H5222 | HEK293 | Human Fc gamma RIIIB / CD16b (NA2) Protein, His Tag (SPR & BLI & MALS verified) |  |
| CDB-H5294 | HEK293 | Human Fc gamma RIIIB / CD16b (NA2) Protein, SUMO,His Tag (MALS & BLI-verified) |  | |
| CDB-H82Ea | HEK293 | Biotinylated Human Fc gamma RIIIB / CD16b (NA2) Protein, His,(SPR & BLI & MALS verified) |  | |
| Fc gamma RIII / CD16 | CDA-M52H8 | HEK293 | Mouse Fc gamma RIII / CD16 Protein, His Tag (SPR & BLI & MALS verified) |  |
| FC6-M82E0 | HEK293 | Biotinylated Mouse Fc gamma RIII / CD16 Protein, His,(MALS & SPR verified) |  | |
| FC6-C52H9 | HEK293 | Cynomolgus Fc gamma RIII / CD16 Protein, His Tag (MALS & BLI verified) |  | |
| FC6-C82E0 | HEK293 | Biotinylated Cynomolgus Fc gamma RIII / CD16 Protein, His,(MALS & BLI verified) |  | |
| FC6-R52H6 | HEK293 | Rhesus macaque Fc gamma RIII / CD16 Protein, His Tag (SPR & BLI & MALS verified) |  | |
| Fc gamma RIV / CD16-2 | FC4-M52H3 | HEK293 | Mouse Fc gamma RIV / CD16-2 Protein, His Tag (MALS verified) |  |
| FC4-M82E8 | HEK293 | Biotinylated Mouse Fc gamma RIV / CD16-2 Protein, His,(BLI verified) |  |
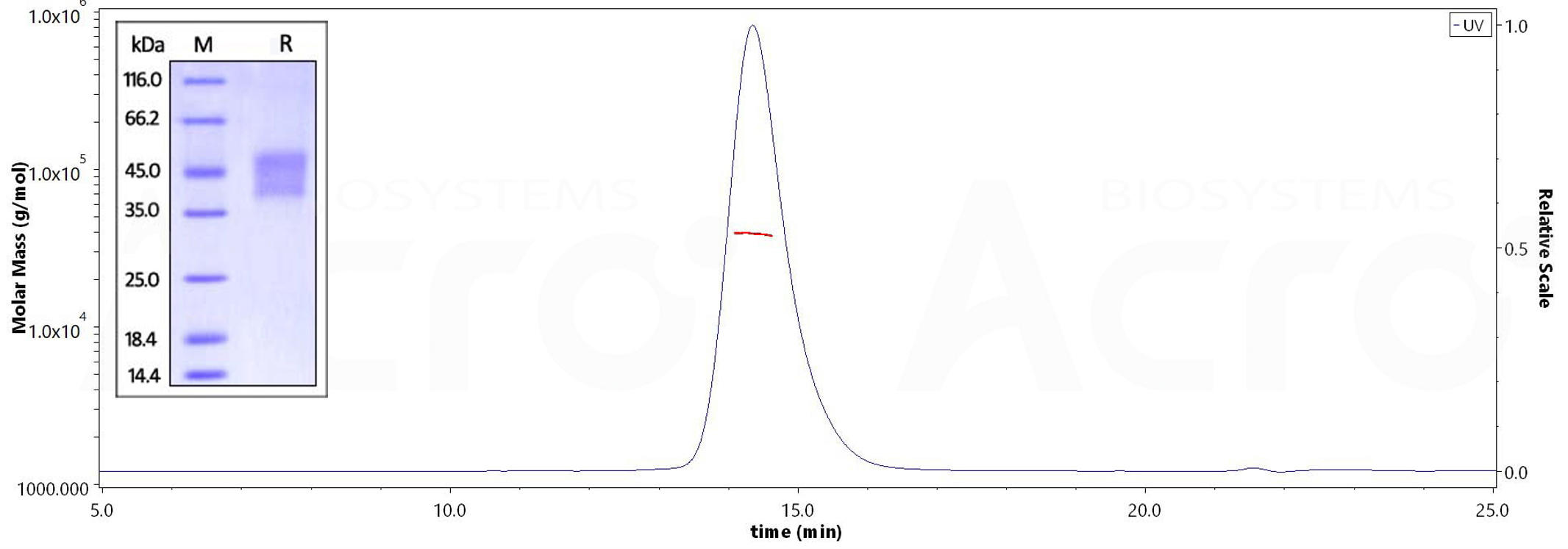
Human CD16a (V176), His Tag on SDS-PAGE under reducing (R) condition. The gel was stained overnight with Coomassie Blue. The purity of the protein is greater than 95%. The purity of Human CD16a (V176), His Tag (Cat. No. CD8-H52H4) is more than 95% and the molecular weight of this protein is around 35-45 kDa verified by SEC-MALS.
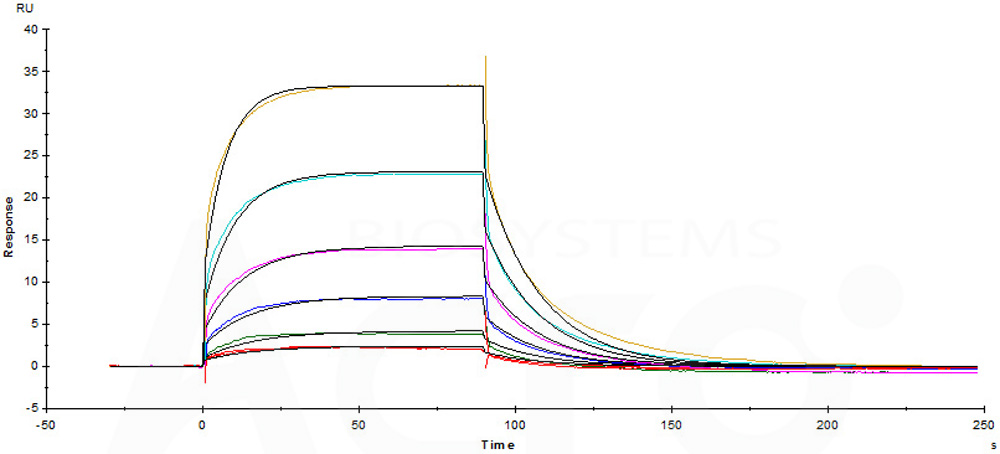
Immobilized Biotinylated Human CD16a (F176), Avitag, His Tag (Cat. No. CDA-H82E8) on SA Chip can bind Rituximab with an affinity constant of 1.33 μM as determined in a SPR assay (Biacore T200).
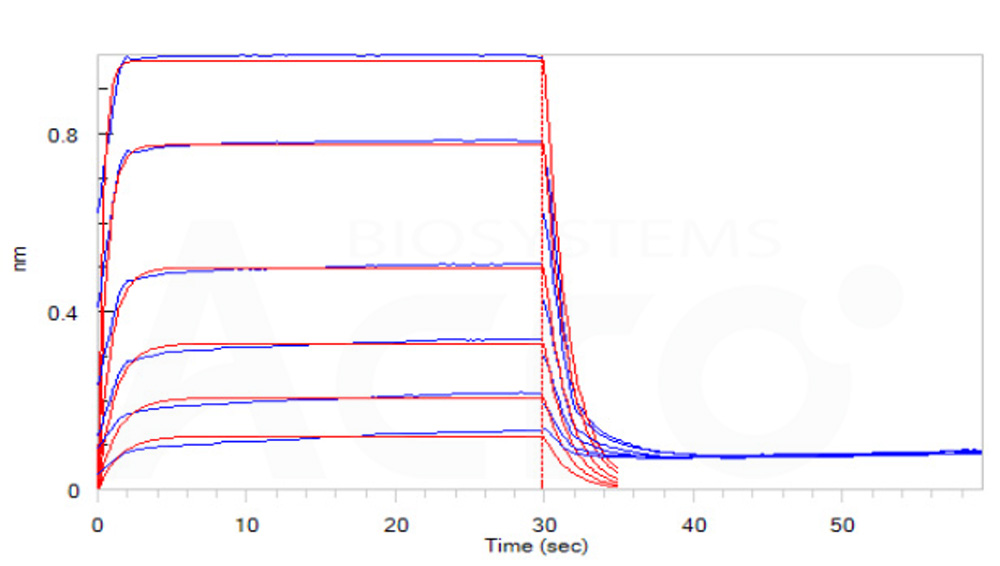
Loaded Biotinylated Human CD16b (NA1), His, Avitag (Cat. No. CDB-H82E4) on SA Biosensor, can bind Rituximab with an affinity constant of 7.7 μM as determined in BLI assay (ForteBio Octet Red96e)
1. Wang, F., Lau, J.K.C. & Yu, J. The role of natural killer cell in gastrointestinal cancer: killer or helper. Oncogene. 2021, 40: 717–730. https://doi.org/10.1038/s41388-020-01561-z
2. Hyun Don Yun, Martin Felices, et al. Trispecific killer engager CD16xIL15xCD33 potently induces NK cell activation and cytotoxicity against neoplastic mast cells. Blood Adv. 2018, 2(13): 1580–1584. doi: 10.1182/bloodadvances.2018018176
3. Karie Runcie, Daniel R. Budman,et al. Bi-specific and tri-specific antibodies- the next big thing in solid tumor therapeutics. Molecular Medicine. 2018, 24:50. https://doi.org/10.1186/s10020-018-0051-4
4. Jingjing Wu, Jiaping Fu, et al. AFM13: a first-in-class tetravalent bispecific anti-CD30/CD16A antibody for NK cell-mediated immunotherapy. Journal of Hematology & Oncology. 2015, 8:96. DOI 10.1186/s13045-015-0188-3
5. Uwe reusch, Carmen Burkhardt, et al. A novel tetravalent bispecific TandAb(CD30/CD16A) efficiently recruits NK cells for the lysis of CD30+ tumor cells. mAbs, 2014, 6:3, 728–739. DOI: 10.4161/mabs.28591
6. Susanne Wingert, Uwe Reusch, et al. Preclinical evaluation of AFM24, a novel CD16A specific innate immune cell engager targeting EGFR-positive tumors. mAbs, 2021, 13:1. DOI: 10.1080/19420862.2021.1950264
7. Martin Felices1, Todd R. Lenvik, et al. Potent Cytolytic Activity and Specific IL15 Delivery in a 2nd Generation Trispecific Killer Engager. Cancer Immunol Res. 2020, 8(9): 1139–1149. doi:10.1158/2326-6066. CIR-19-0837
8. Antibody Drug Discovery Downloaded from www.worldscientific.com by 146.151.196.179 on 10/03/14
This web search service is supported by Google Inc.
